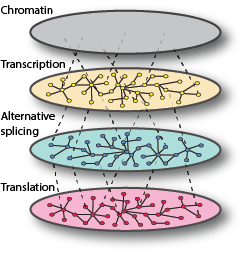
Changes in gene expression have significant impacts on biological phenotypes, including human disease. These changes can occur in all gene expression processes, including mRNA transcription, alternative splicing, translation into protein, and mRNA and protein turnover. The regulation of these processes involves complex networks comprised of cis-acting DNA and RNA sequence elements, and trans-acting factors. Over the past decade it has become increasingly clear that changes in both types of network components have contributed to widespread differences in mRNA abundance, both among and between species. However, species differences in mRNA abundance may not accurately reflect differences in protein expression and comparatively little is known regarding the evolution of post-transcriptional gene regulatory processes. We are currently using both laboratory and computational based methods to investigate upstream open reading frames (uORFs), and how they affect translation regulatory divergence in multiple Saccharomyces species.